Our research combines experimental approaches with computational methods to identify functionally related genes, i.e. gene modules, that work together to express traits, such as metabolites, biopolymers, tissues, and cell types. Identification of gene modules is the first step in gene function prediction needed to improve and utilize the traits.
Our current focus in gene function prediction is the elucidation of biosynthetic pathways that produce valuable secondary metabolites in medicinal plants. Knowing the enzymes in a metabolic pathway is a prerequisite for an efficient large-scale production of these metabolites for use in medicine and industry. Our second focus in gene function prediction is the evolution of adaptation to environmental stresses. To understand how stress-related traits evolve, we are analyzing the development of these modules in the plant kingdom. Finally, to make our findings available to the public, we generate online databases that provide user-friendly tools to browse our networks and predictions.
Our current focus in gene function prediction is the elucidation of biosynthetic pathways that produce valuable secondary metabolites in medicinal plants. Knowing the enzymes in a metabolic pathway is a prerequisite for an efficient large-scale production of these metabolites for use in medicine and industry. Our second focus in gene function prediction is the evolution of adaptation to environmental stresses. To understand how stress-related traits evolve, we are analyzing the development of these modules in the plant kingdom. Finally, to make our findings available to the public, we generate online databases that provide user-friendly tools to browse our networks and predictions.
|
Gene Modules
|
Secondary Metabolites
|
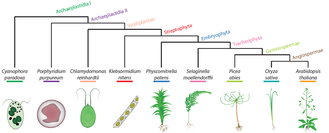
Plant Kingdom
|